Our focus is on the investigation of human pathogens of bacterial, fungal and viral origin including: how pathogens interact with their host, innovative ways of identifying and diagnosing infection, and novel infection control strategies using contemporary antimicrobial treatments. These studies encompass pathogens that affect a variety of different organ systems, host immune responses, and the development of both synthetic and natural antimicrobial treatments.
Please follow on Twitter: @MIRG_CMU
Research / Innovation Areas
Host-pathogen interactions
These studies use in vitro and in vivo (Galleria mellonella) infection models to understand chronic wound infection, sexually transmitted infections, respiratory tract infections, persistent viral infection, and other clinically relevant infections.
Rebecca Aicheler’s work aims to understand how human cytomegalovirus (HCMV) modulates the natural killer (NK) cell response. HCMV is a risk to unborn foetuses and a major cause of congenital malformation. Lauren Jones, a PhD student working within group is currently investigating the role of HCMV-encoded IL-10 (v-IL-10) in kidney transplant patients
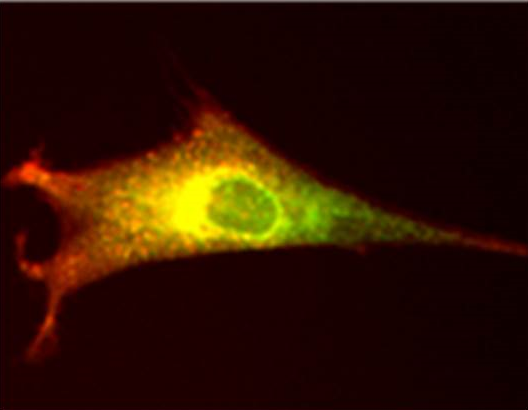
Human cytomegalovirus UL141 prevents NK cells from binding to CD155 and the NK cell is inhibited.
Mike Beeton’s research focuses on understanding the role mycoplasmas and ureaplasmas play in human health and disease, characterising their mechanisms of antibiotic resistance, understanding their role in preterm birth and respiratory tract infections.
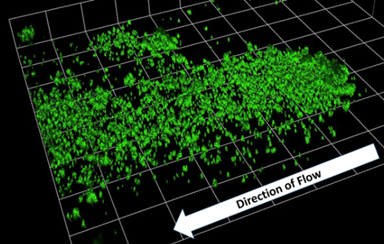
Biofilm formation by Ureaplasma parvum when grown under flow conditions.
Sarah Maddocks investigates interactions between pathogens in co-cultured biofilms using an in vitro chronic wound biofilm model. This work aims to understand differential colonisation, altered virulence, and the impact these have on progression of infection. Sarah also studies the role of small colony variant bacteria including how they persist in chronic infection.
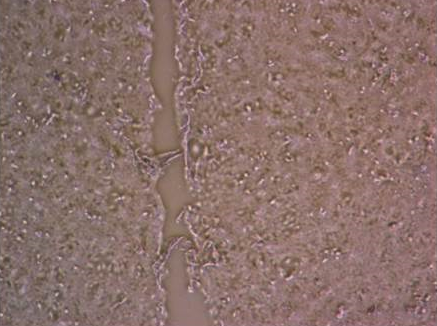
Impaired healing of human keratinocytes exposed to Staphylococcus aureus.
Novel antimicrobials treatments and disinfectants
This research investigates natural and synthetic approaches to address the ongoing problem of antimicrobial resistance.
Mike Beeton is investigating the use of bacteriophage isolated from the environment to tackle infection caused by clinically relevant pathogens. These studies have expanded to include the design of targeted interventions against mycoplasmas and ureaplasmas.
James Blaxland’s research focuses on the use of ozone to decontaminate the bacterium Listeria monocytogenes from both food and food preparation areas. Working with a number of ozone device manufacturers, James is identifying a standard method for determining ozone efficacy within a range of environments. James is also screening and characterising a range of natural materials including honey and hops to determine their antimicrobial potential. His research interests span both clinical and food microbiology.

Paul Livingstone’s work aims to identify predatory microorganisms with untapped biosynthetic potential. Such microorganisms consume prey microbes through the secretion of antimicrobial substances, and could be used to combat numerous clinically relevant pathogens.
Sarah Maddocks works with Mike to investigate the use of bacteriophage to disrupt polymicrobial biofilm in combination with systemic antibiotics, in an in vitro chronic wound biofilm model. This model is similarly used to test novel topical antimicrobial dressings and gels.

Testing antimicrobials in an in vitro chronic wound biofilm model.
Microbial Genomics and Taxonomy
Paul Livingstone uses next generation sequencing and contemporary genomic analysis to identify and characterise environmental bacteria with metabolic potential that could be exploited for the degradation of pollutants. Paul has also identified new taxa of predatory microorganisms of various genera and is investigating the genomic basis for predation.
Paul is also working with Sarah to understand the genomic basis of the small colony variant phenotype, as well as metagenomic analysis of chronic infected wounds.
Group members
 |
 |
 |
Senior Lecturer in
Immunology | Senior Lecturer in
Medical Microbiology | Dr James Blaxland,
Lecturer in Food
Microbiology |
 |

|  |
Dr Paul Livingstone, Lecturer in Medical Microbiology |
Dr Sarah Maddocks,
Senior Lecturer in
Microbiology (Group Lead)
| Lauren Jones,
PhD Student
|

| 
| |
Recent Publications from MIRG
Heymans C., Heij L.R., Lenaerts K., Dulk M., Hadfoune M., Heugten C., Spiller O.B., Beeton M.L., Stock S. J., Jobe A. H., Payne M.S., Kemp M.W., Kramer B.W., Plat J., van Gemert W. and Wolfs T.G.A.M. (2020). Prophylactic Intra-Uterine β-Cyclodextrin Administration during Intra-Uterine Ureaplasma parvum Infection Partly Prevents Liver Inflammation without Interfering with the Enterohepatic Circulation of the Fetal Sheep.
Heymans C., de Lange I.H., Hütten M.C., Lenaerts K., de Ruijter N.J.E., Kessels L.C.G.A., Rademakers G., Melotte V., Boesmans W., Saito M., Usuda H., Stock S.J., Spiller O.B., Beeton M.L., Payne M.S., Kramer B.W., Newnham J.P, Jobe A.H., Kemp M.W., van Gemert W.G. and Wolfs T.G.A.M. (2020) . Chronic intra-uterine Ureaplasma parvum infection induces injury of the enteric nervous system in ovine fetuses
Zwarycz, A. S., Livingstone, P. G. and Whitworth D. E. (2020). Within-species variation in OMV cargo proteins: the Myxococcus xanthusOMV pan-proteome.
Anbarasu A, Ramaiah S & Livingstone P.G (2020). Vaccine repurposing approach for preventing COVID 19: can MMR vaccines reduce morbidity and mortality?
Saracato, M and Blaxland, J (2020) Dairy-Free Imitation Cheese: Is further Development Required?
Anastasiia Mykhailenko Andriy Utevsky Olexii Solodiankin Oksana Zlenko Olha Maiboroda Vitaliy Bolotin James Blaxland Anton Gerilovych (2020). First record of Serratia marcescens from Adelie and Gentoo penguin faeces collected in the Wilhelm Achipelago, Graham Land, West Antartica
Dua, K., Blaxland, J., Gulati, M., (2020). Metabiotics in Colorectal Cancer: Crosstalk between Gut Microbiota and Host Pathology' Book Chapter in: "An Update on Probiotic Research in Therapeutics"
Virginia-Maria Vlahava1*, Isa Murrell1*, Lihui Zhuang1, Rebecca J. Aicheler2, Eleanor Lim3, Kelly L. Miners1, Kristin Ladell1, Nicolás M. Suárez4, David A. Price1, Andrew J. Davison4, Gavin W.G. Wilkinson1, Mark R. Wills3, Michael P. Weekes5, Eddie C.Y. Wang1, Richard J. Stanton1§ Monoclonal antibodies targeting non-structural viral antigens can activate ADCC against human cytomegalovirus.Journal of Clinical Investigation. In press
Heath N., Rowlands R., Webster G., Mahenthiralingam E. and Beeton M. (2020). Antimicrobial activity of enacyloxin IIa and gladiolin against the urogenital pathogens Neisseria gonorrhoeae and Ureaplasma spp
James Chambers 1, Natalie Sparks 1, Natashia Sydney 1, Paul G Livingstone 1 2, Alan R Cookson 1, David E Whitworth 1 Comparative genomics and pan-genomics of the Myxococcaceae, including a description of five novel species: Myxococcus eversor sp. nov., Myxococcus llanfairpwllgwyngyllgogerychwyrndrobwllllantysiliogogogochensis sp. nov., Myxococcus vastator sp. nov., Pyxidicoccus caerfyrddinensis sp. nov. and Pyxidicoccus trucidator sp. nov
Duckworth P. F., Maddocks, S. E., Rahatekar, S. S., Barbour M. E (2020) Alginate films augmented with chlorhexidine hexametaphosphate particles provide sustained antimicrobial properties for application in wound care. Journal of Material Science. 31: 33
Livingstone, P., Ingleby, O., Girdwood, S., Cookson, A., Morphew, R., Whitworth, D. (2019)
Predatory organisms with untapped biosynthetic potential. A description of eight novel Corallococcus species : Corallococcus aberystwythiensis sp. nov., Corallococcus carmarthensis sp. nov., Corallococcus exercitus sp. nov., Corallococcus interemptor sp. nov., Corallococcus llansteffanensis sp. nov., Corallococcus praedator sp. nov., Corallococcus sicarius sp. nov., and Corallococcus terminator sp. nov.
DOI:10.1128/AEM.01931-19 Applied and Environmental Microbiology
Sutton, D., Livingstone, P., Furness, E., Swain, M., Whitworth, D (2019) Genome-Wide Identification of Myxobacterial Predation Genes and Demonstration of Formaldehyde Secretion as a Potentially Predation-Resistant Trait of Pseudomonas aeruginosa DOI:10.3389/fmicb.2019.0265 Frontiers in Microbiology
Beeton M.L, Zhang X.S, Uldum S. A, Bébéar C, Dumke R, Gullsby K, Ieven M, Loens K, Nir-Paz R, Pereyre S, Spiller O.B, Chalker VJ and the ESCMID Study Group for Mycoplasma and Chlamydia Infections (ESGMAC) Mycoplasma pneumoniae sub-group. Mycoplasma pneumoniae infections across Europe and Israel (2011-2016). Eurosurveillance. 2019. In Press
Beeton ML, Payne MS and Jones LC. The role of Ureaplasma spp. in the development of non-gonococcal urethritis and infertility among men. Clin. Microbiol. Rev. 2019. Jul 3;32(4). pii: e00137-18. doi: 10.1128/CMR.00137-18
van Gorp C, de Lange IH, Spiller OB, Dewez F, Cillero Pastor B, Heeren RMA, Kessels L, Kloosterboer N, van Gemert WG, Beeton ML, Stock SJ, Jobe AH, Payne MS, Kemp MW, Zimmermann LJ, Kramer BW, Plat J and Wolfs TGAM. Protection of the Ovine Fetal Gut against Ureaplasma-Induced Chorioamnionitis: A Potential Role for Plant Sterols. Nutrients. 2019 Apr 27;11(5). doi: 10.3390/nu11050968.
Duckworth, P. F., Rowlands, R. S., Barbour, M. E. and Maddocks S. E. (2018) A novel flow system to establish experimental biofilms for modelling chronic wound infection and testing the efficacy of wound dressings. Microbiological Research. 215: 141-147.
Alves, P.M., Al-Badi, E., Withycombe, C., Jones, P.M., Purdy, K.J., Maddocks, S.E. (2018). Interaction between Staphylococcus aureus and Pseudomonas aeruginosa is beneficial for colonisation and pathogenicity in a mixed-biofilm. Pathogens and Disease doi: 10.1093/femspd/fty003
Livingstone, P., Millard, A., Swain, M. T. & Whitworth, D. E. (2018) Transcriptional changes when Myxococcus xanthus preys on Escherichia coli suggest myxobacterial predators are consititutively toxic Microbial Genomics. 152
Barbour, M.E., Maddocks, S.E., Grady, H.J., Roper, J.A., Bass, M.D., Collins, A.M., Dommett, R.M. and Saunders, M.(2016) Chlorhexidine hexametaphosphate as a wound care material coating: antimicrobial efficacy, toxicity and effect on healing. Nanomedicine. 11:2049-57
Fielding, C.A., Weekes, M.P., Nobre, L.V., Ruckova, E., Wilkie, G.S., Paulo, J.A., Chang, C., Suárez, N.M., Davies, J.A., Antrobus, R., Stanton, R.J., Aicheler, R.J., Nichols, H., Vojtesek, B., Trowsdale, J., Davison, A.J., Gygi, S.P., Tomasec, P., Lehner, P.J., Wilkinson, G.W. (2017). Control of immune ligands by members of a cytomegalovirus gene expansion suppresses natural killer cell activation. Elife. doi: 10.7554/eLife.22206.
Eissa, A., Blaxland, J., Williams, R., Metwally, K., El-Adl, S., Lashine, E., Baillie, L., Simons, C. (2016) Adenine and benzimidazole-based mimics of REP-3123 as antibacterial agents against Clostridium difficile and Bacillus anthracis: Design, synthesis and biological evaluation DOI: 10.1016/j.bfopcu.2016.06.002 Bulletin of Faculty of Pharmacy, Cairo University
Weekes, M.P., Tomasec, P., Huttlin, E.L., Fielding, C.A., Nusinow, D., Stanton, R.J., Wang, E.C., Aicheler, R.J., Murrell, I., Wilkinson, G.W., Lehner, P.J., Gygi, S.P. (2014). Quantitative temporal viromics: an approach to investigate host-pathogen interaction. Cell. 157(6):1460-72. doi: 10.1016/j.cell.2014.04.028.
*Smith .W., *Tomasec. P., *Aicheler, R.J., Loewendorf, A., Nemčovičová, I., Wang, E.C., Stanton, R.J., Macauley, M., Norris, P., Willen, L., Ruckova, E., Nomoto, A., Schneider, P., Hahn, G., Zajonc, D.M., Ware, C.F., Wilkinson, G.W., Benedict, C.A. (2013). Human cytomegalovirus glycoprotein UL141 targets the TRAIL death receptors to thwart host innate antiviral defenses. Cell Host Microbe.13(3):324-35. doi: 10.1016/j.chom.2013.02.003.
* Joint first authors
Research Students:
Ammara Khalid (PhD student) is modelling complex multi-species biofilms in a dynamic chronic wound system with the aim of understanding interspecies interactions and the impact they have on disease progression. Ammara is also investigating how bacterial interactions can be exploited for the effective management of chronic wounds.
Benita Arakal (PhD student): Bioprospecting Soil-borne Myxobacterial Isolates and Assessing its Predatory Effects on a Myriad of Drug-resistant Infection Models.
Lauren Jones (PhD student): Human Cytomegalovirus (HCMV) Encoded Interleukin-10 (vIL-10): Investigating the Immunomodulatory Effects of vIL-10 and its Suitability as a Novel Marker of Viral Reactivation in Kidney Transplant Patients. Supported by KESS 2 and Kidney Wales Foundation. Supervisors: Dr. Rebecca Aicheler, Dr. Sarah Maddocks and Professor Ian Humphreys (Cardiff University)
Robert Foxwell (PhD student): Assessing the potential of nanoparticle-based bee supplements as a means of reducing the frequency of colony collapse in Apis mellifera hives and the impact that these supplements have on hive health, hive population dynamics, hive honey, pathogen and parasitic infestation occurrence. Supported by BBI Solutions. Supervisors: Dr. James Blaxland and Dr. Sarah Maddocks
Andreea-Gabriela Nedelea (MRes student): Optimising an ex vivo porcine skin model for chronic wound diagnosis and treatment. Supervisors: Dr. Sarah Maddocks, Dr Lori Robins (University of Washington, Bothell, USA)
Thomas Burden (MRes student): Investigating how small colony variant bacteria evade immune detection. Supervisors: Dr Rebecca Aicheler, Dr Sarah Maddocks
Collaborators
Dr Ambikesh Jayal, Cardiff Metropolitan University
Mr Gareth Thomas, Cardiff Metropolitan University
Dr Paul Smith, Cardiff Metropolitan University
Dr Brad Spiller, Cardiff University
Dr Gavin Wilkinson, Cardiff University
Dr Richard Stanton, Cardiff University
Dr Matthew Bates, MCBA Consulting Ltd.
Dr Hajer Taleb, Neem Biotech
Dr Robert Atterbury, University of Nottingham
Dr Vicki Chalker, Public Health England
Dr Nicholas Tucker, University of Strathclyde
Dr Kevin Purdy, University of Warwick
Prof Keith Harding, Welsh Wound Innovation Centre
Dr Matt Payne, University of Western Australia
Dr Lucy Furfaro, University of Western Australia
Dr Steven Coles, University of Worcester
Dr Lori Robins, University of Washington
Professor Ian Humphreys, Cardiff
Examples of Funding
Cardiff Metropolitan Research and Innovation Services
Institute of Biomedical Sciences
KESS2
Microbiology Society
Sir Halley Stewart Trust
Society for Applied Microbiology
The Royal Society
